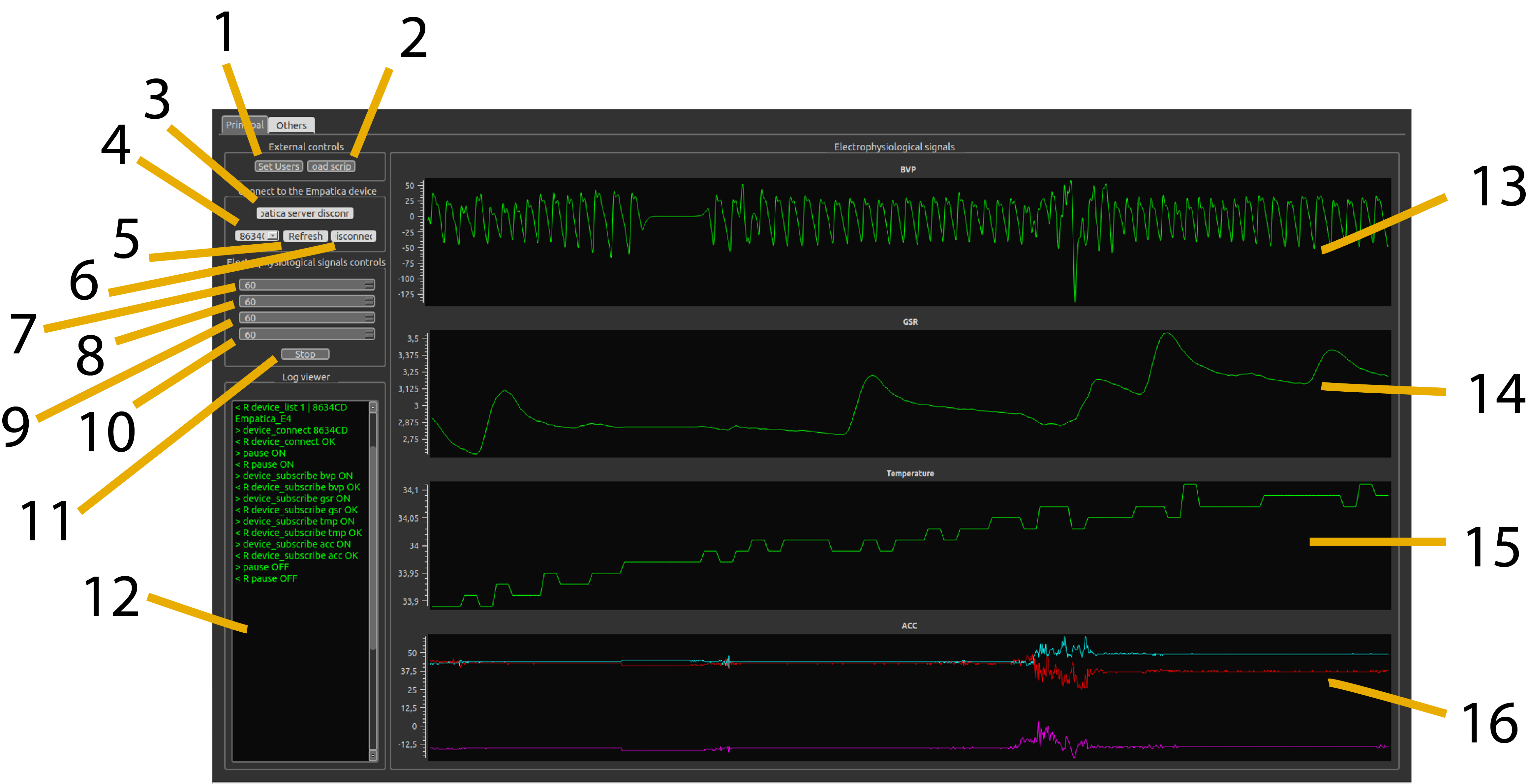
BIOSIGNALS software interface: Real-time monitoring of BVP, GSR, Temperature, and Accelerometer data with PyQt5 GUI
The Challenge: Synchronised Multi-Modal Physiological Data in Real-Time
In affective computing and human-robot interaction research, understanding human emotions requires capturing multiple physiological signals simultaneously whilst maintaining precise temporal synchronisation with experimental events. However, most commercial solutions are either prohibitively expensive, closed-source, or lack the sub-100ms event synchronisation needed for dynamic HRI protocols.
During my PhD research on Emotional Human-Robot Interaction at Universidad Nacional de Educación a Distancia (UNED), I encountered a fundamental technical gap: no existing system could acquire heterogeneous physiological signals at different sampling rates whilst providing TCP/IP-based remote triggering for closed-loop robotic control.
The Multi-Modal Synchronisation Problem
Physiological signals operate at vastly different temporal scales:
- BVP (Blood Volume Pulse): 64 Hz — cardiac cycle requires high temporal resolution
- GSR (Galvanic Skin Response): 4 Hz — slow sympathetic responses
- TMP (Temperature): 4 Hz — even slower thermal changes
- ACC (3-axis Accelerometer): 32 Hz — motion artifact detection
Challenge: Synchronise these heterogeneous streams with external events (robot actions, stimuli) at <50ms latency whilst maintaining thread-safe concurrent processing.
The system needed to:
- Acquire 4 signal modalities simultaneously with per-channel thread-safe ring buffers
- Visualise in real-time (>30 FPS) without blocking data acquisition threads
- Respond to TCP/IP triggers with minimal latency for closed-loop HRI experiments
- Annotate events with microsecond timestamps embedded in EDF+ format
- Process HRV and GSR features online for real-time emotion classification
- Maintain modular architecture for integration with external ML pipelines
Research Context: Affective Robot Storytelling
BIOSIGNALS was developed as part of the "Emotional Human-Robot Interaction with Physiological Signals" doctoral project at UNED's AI Department. The system enabled experiments where NAO robots adapted narrative delivery based on children's real-time emotional states detected through synchronised BVP, GSR, and facial expression analysis.
Key achievement: 74% accuracy in dynamic emotion classification by fusing HRV features (valence) with GSR arousal detection, enabling truly adaptive social robotics.
System Architecture: Multi-Threaded Event-Driven Design
BIOSIGNALS implements a multi-threaded event-driven architecture with strict separation of concerns to handle concurrent I/O, real-time visualisation, and remote control without race conditions.
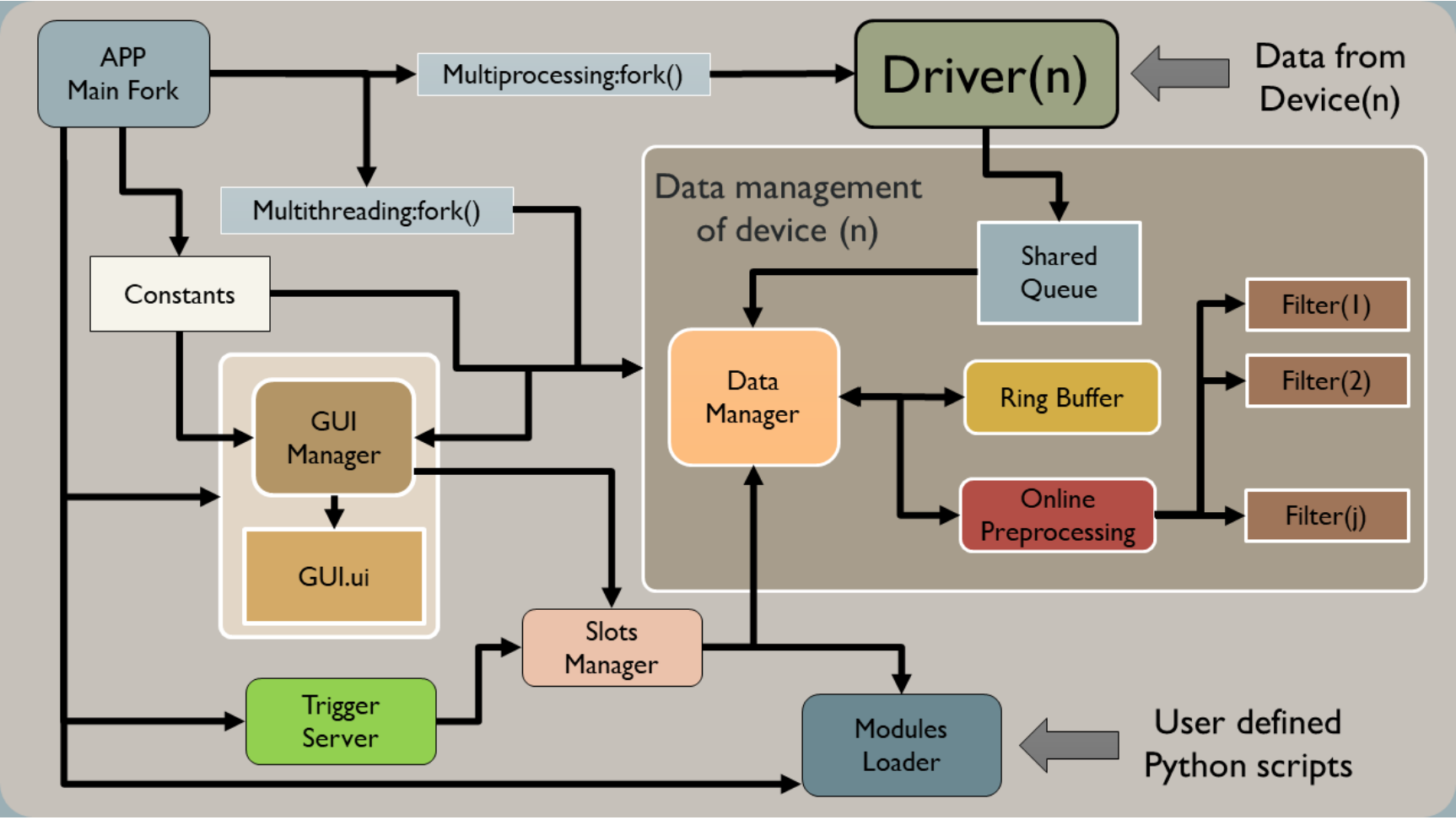
Software architecture: Multi-threaded design with state machine, data managers, ring buffers, and TCP/IP trigger server
Architecture Overview
┌──────────────────────────────────────────────────────────────┐
│ BIOSIGNALS_APP_01.py (Main Thread) │
│ │
│ ┌──────────────┐ ┌──────────────┐ ┌──────────────┐ │
│ │ State │─▶│ Socket │─▶│Data Managers │ │
│ │ Machine │ │ Threads │ │ (x4 types) │ │
│ └──────────────┘ └──────────────┘ └──────────────┘ │
│ │ │ │ │
│ ▼ ▼ ▼ │
│ ┌──────────────┐ ┌──────────────┐ ┌──────────────┐ │
│ │ GUI Layer │ │ Trigger │ │Ring Buffers │ │
│ │ (PyQt5) │ │ Server │ │(Thread-Safe) │ │
│ └──────────────┘ └──────────────┘ └──────────────┘ │
└──────────────────────────────────────────────────────────────┘
1. Finite State Machine for Lifecycle Management
The application implements an FSM to manage connection states and recording transitions:
class MyApp(QtWidgets.QApplication):
"""
Main application with state machine control.
States:
SERVER → Disconnected from Empatica Server
DEVICE → Connected, waiting for E4 device
VIEW → Device streaming data
Substates (in VIEW):
OFF → Paused
ON → Recording
WAIT_PAUSE → Transitioning
"""
def __init__(self):
self.state = "SERVER"
self.substate = ""
self.trigger_control = False # Remote trigger flag
self.pause_control = True # Local pause flag
@QtCore.pyqtSlot(str)
def trigger_event(self, action):
"""Handle TCP/IP trigger events with FSM logic"""
if self.state == "VIEW":
if not self.trigger_control: # Not recording
if action == 'start':
# Create EDF files
for dmg in self.dmgs:
dmg.create_file()
self.start()
self.thread.flag.set()
self.trigger_control = True
else: # Already recording
if action in ['start', 'stop']:
# Stop and save
self.start()
for dmg in self.dmgs:
dmg.save_streamData()
dmg.reset_data_store()
self.trigger_control = False
else:
# Annotate event during recording
for dmg in self.dmgs:
dmg.online_annotation(action)
2. Thread-Safe Ring Buffer Implementation
Each signal has its own circular buffer with mutex-protected concurrent access:
class RingBuffer(QtCore.QThread):
emitter = QtCore.pyqtSignal()
def __init__(self, channels, num_samples, sample_rate):
self.max = num_samples
self.data = np.zeros((self.max, channels))
self.cur = 0 # Current write position
self.cur_show = self.max # Display offset
# Visualisation control
self.seconds = 6
self.control = sample_rate * self.seconds
def append(self, x):
"""O(1) circular insertion"""
self.cur = self.cur % self.max
self.data[self.cur, :] = np.array(x)
self.cur += 1
if self.cur_show > 0:
self.cur_show -= 1
# Emit signal every N seconds for GUI update
if (self.cur_show == 0) and ((self.cur % self.control) == 0):
self.emitter.emit()
def get(self):
"""Return ordered data (oldest → newest)"""
data = np.vstack((self.data[self.cur:, :],
self.data[:self.cur, :]))
return data[self.cur_show:, :]
Advantages
- ⚡ O(1) complexity for insert/retrieve
- 🔒 Thread-safe with mutex locks
- 📊 Dynamic windows without reallocation
- 🔔 Async notifications via Qt signals
Performance Metrics
- Buffer insertion: <1 ms
- GUI update (64 Hz): 15.6 ms (real-time)
- TCP trigger latency: <50 ms (LAN)
- Timestamp precision: 1 ms
3. TCP/IP Trigger Server for Remote Control
A dedicated thread handles remote commands for experimental automation:
class TriggerServer(QtCore.QThread):
socket_emitter = QtCore.pyqtSignal(str)
def create_socket(self):
self.sock = socket.socket(socket.AF_INET, socket.SOCK_STREAM)
self.server_address = ('localhost', 10000)
self.sock.bind(self.server_address)
self.activated = True
def run(self):
self.sock.listen(1)
while self.activated:
connection, client = self.sock.accept()
while True:
data = connection.recv(128)
if data:
# Emit event to main thread
self.socket_emitter.emit(data.decode())
# Client usage (from any Python script)
from COM.trigger_client import trigger_client
tc = trigger_client('192.168.1.100', 10000)
tc.create_socket()
tc.connect()
tc.send_msg(b'start') # Begin recording
tc.send_msg(b'stimulus_happy') # Annotate event
tc.send_msg(b'stop') # Stop and save
Heart Rate Variability: Robust Multi-Stage Peak Detection
Extracting clean NN intervals (Normal-to-Normal) from photoplethysmography signals is challenging due to motion artifacts, low SNR, and inter-individual morphology variations. Our pipeline implements a 6-stage robust algorithm:
HRV Processing Pipeline
- Signal Inversion & Savitzky-Golay Filtering: Denoise whilst preserving peak sharpness
- Adaptive Upper Envelope: Sliding window maximum for baseline tracking
- Peak Enhancement via Sigmoid Transform: Amplify peaks relative to dynamic baseline
- Peak Detection with Prominence Filtering: scipy.signal.find_peaks with physiological constraints
- Physiological Range Filter: Reject HR <50 or >120 bpm (arrhythmia detection)
- Convolutional STD Artifact Detection: O(n) outlier removal using kernel convolution
def compute_nni(hrdata, sample_rate=64, sliding_window=0.5,
prominence=0.1, dist_q1=50, dist_q2=120,
std_window=6, std_th=130, method='remove'):
"""
Compute NN intervals from BVP signal with robust artifact handling.
Args:
hrdata: Raw BVP signal (numpy array)
sample_rate: Sampling frequency (Hz)
sliding_window: Window for adaptive envelope (seconds)
prominence: Minimum peak prominence (normalized)
dist_q1, dist_q2: Valid HR range (bpm) [50-120]
std_window: STD convolution window (samples)
std_th: Artifact threshold (STD threshold)
method: 'remove', 'iqr', 'modified_z' for artifact handling
Returns:
nni_revised: Clean NN intervals (milliseconds)
"""
# Step 1: Signal inversion and smoothing
hrdata_inv = hrdata * (-1)
roll_mean = savgol_filter(hrdata_inv, 81, 2) # Order-2 S-G
# Step 2: Adaptive upper envelope
windowsize = int(sliding_window * sample_rate)
add = np.zeros(int(windowsize / 2))
add[:] = np.nan
hrdata_ext = np.concatenate((add, hrdata_inv, add))
roll_max = []
for i in range(len(hrdata)):
roll_max.append(np.nanmax(hrdata_ext[i:i+windowsize]))
sroll_max = savgol_filter(roll_max, 51, 2)
mn = 0.3 * np.std(sroll_max)
sroll_max = sroll_max + mn
# Step 3: Peak enhancement (simplified representation)
simpleHR_1 = (hrdata_inv - roll_mean) * (hrdata_inv > roll_mean)
envoltorio = minmax(sroll_max - roll_mean)
simpleHR_2_raw = sigmoid(0, 2, 5, envoltorio) * simpleHR_1
simpleHR_2 = savgol_filter(simpleHR_2_raw, 31, 2)
# Step 4: Peak detection
peaksx = np.where((simpleHR_2 > 0))[0]
peaksy = simpleHR_2[peaksx]
peaks, _ = find_peaks(peaksy, prominence=prominence)
# Step 5: NN intervals + physiological filter
nni = tools.nn_intervals((peaksx[peaks] / sample_rate) * 1000)
hr = tools.heart_rate(nni)
index = np.logical_and((hr >= dist_q1), (hr <= dist_q2))
nni_revised = nni[index]
# Step 6: Convolutional STD artifact detection (O(n))
std = std_convoluted(nni_revised, std_window)
index_std = [i for i in range(len(std)) if std[i] > std_th]
# Remove artifact segments
if method == 'remove':
nni_revised[index_std] = np.nan
elif method == 'iqr':
nni_revised[index_std] = outliers_iqr_method(nni_revised)
return nni_revised[~np.isnan(nni_revised)]
def std_convoluted(nni, N):
"""
Compute local STD via convolution (O(n) vs O(n*k) sliding window).
Uses: Var(X) = E[X²] - E[X]²
Convolves both X and X² with uniform kernel.
"""
im = np.array(nni, dtype=np.float64)
im2 = im**2
kernel = np.ones(2*N+1)
s = convolve(im, kernel, mode="same") # Sum of X
s2 = convolve(im2, kernel, mode="same") # Sum of X²
ns = convolve(np.ones(im.shape), kernel, mode="same")
return np.sqrt((s2 - s**2 / ns) / ns) # STD formula
Key Innovation: O(n) Artifact Detection
Traditional sliding-window STD computation is O(n*k). By using kernel convolution to compute E[X²] and E[X]² simultaneously, we achieve O(n) complexity with FFT-based convolution—critical for real-time processing of long recordings.
HRV Feature Extraction: Temporal, Frequency, Non-Linear
Once clean NN intervals are obtained, we extract a comprehensive feature set spanning three domains:
Time-Domain
- SDNN: Global variability
- RMSSD: Vagal tone (parasympathetic)
- pNN50: % of NNI differing >50ms
- Mean HR: Average heart rate
Frequency-Domain (Welch PSD)
- VLF: 0.003-0.04 Hz
- LF: 0.04-0.15 Hz (sympathetic)
- HF: 0.15-0.4 Hz (parasympathetic)
- LF/HF: Sympatho-vagal balance
Non-Linear
- SampEn: Sample entropy (complexity)
- Poincaré SD1/SD2: Beat-to-beat variability
- DFA α1/α2: Fractal scaling
def compute_features(nni):
"""Extract comprehensive HRV features."""
features = {}
# Temporal domain
features['mean_hr'] = tools.heart_rate(nni).mean()
features['sdnn'] = td.sdnn(nni)[0] # Global variability
features['rmssd'] = td.rmssd(nni)[0] # Vagal tone indicator
features['pnn50'] = td.nn50(nni)[1] # % NNI > 50ms diff
# Frequency domain (Welch PSD)
psd = fd.welch_psd(nni, show=False)
features['hf_lf_ratio'] = psd['fft_ratio'] # Sympatho-vagal
features['lf'] = psd['fft_peak'][1] # 0.04-0.15 Hz
features['hf'] = psd['fft_peak'][2] # 0.15-0.4 Hz
features['log_lf'] = psd['fft_log'][1]
features['log_hf'] = psd['fft_log'][2]
# Non-linear
features['sampen'] = nl.sample_entropy(nni)[0] # Complexity
return features
Physiological Interpretation
- 📈 RMSSD ↑ / HF ↑: High parasympathetic activity → Relaxation, rest-and-digest
- 📉 LF/HF ratio ↑: Sympathetic dominance → Stress, arousal, fight-or-flight
- ⚖️ SDNN ↑: Good autonomic regulation and adaptability
- 🔄 SampEn ↑: Higher complexity → Better cardiovascular health
GSR Decomposition: Tonic and Phasic Components
Galvanic Skin Response (GSR), also known as Electrodermal Activity (EDA), measures skin conductance controlled by sympathetic nervous system activity. The signal comprises two components:
Tonic Component (SCL)
Skin Conductance Level: Slow-varying baseline
- Reflects general arousal state
- Changes over minutes/hours
- Influenced by circadian rhythm, stress
- Features: Mean level, slope, variability
Phasic Component (SCR)
Skin Conductance Response: Rapid event-related peaks
- Response to specific stimuli (1-5s latency)
- Sympathetic activation bursts
- Emotional/cognitive load marker
- Features: Amplitude, frequency, rise/recovery time
Decomposition Algorithm
def extract_gsr_components(gsr_data):
"""
Decompose GSR into tonic and phasic components.
Process:
1. Upsample to 8 Hz (standard for GSR analysis)
2. Rolling mean (window=20) for tonic extraction
3. Subtraction for phasic component
4. Savitzky-Golay refinement of tonic
Args:
gsr_data: DataFrame ['datetime', 'EDA']
Returns:
DataFrame ['EDA', 'phasic', 'tonic']
"""
gsr_data = pd.DataFrame(gsr_data, columns=['datetime', 'EDA'])
sampleRate = 4 # Empatica E4 native rate
startTime = gsr_data.iloc[0, 0]
# Interpolate to 8 Hz
gsr_data = interpolateDataTo8Hz(gsr_data, sampleRate, startTime)
# Tonic: rolling mean
rolling_mean = gsr_data.EDA.rolling(window=20).mean()
# Phasic: signal - tonic
gsr_data['phasic'] = gsr_data.EDA - rolling_mean
# Refine tonic with Savitzky-Golay
window_length = int(len(gsr_data) / 100) * 2 + 1
gsr_data['tonic'] = savgol_filter(gsr_data.EDA,
window_length, 2)
return gsr_data
def compute_phasic_features(gsr_data):
"""
Detect and characterise Skin Conductance Responses (SCRs).
SCR Detection Criteria:
- Amplitude threshold: >0.1 μS
- Must return to baseline
- Minimum 2 inflection points
Features per SCR:
- start/end: Temporal bounds (sample indices)
- peak_locs: Peak location (sample index)
- amp: Amplitude (μS)
- rise_time: Onset → peak (samples)
- recovery_time: Peak → baseline (samples)
Returns:
DataFrame with all detected SCRs
"""
aux1 = np.diff(gsr_data.phasic > 0.1) # Activation
aux2 = np.diff(gsr_data.phasic < 0) # Return
true_list = np.where(aux2)[0]
peaks = {
'start': [], 'end': [], 'peak_locs': [],
'amp': [], 'rise_time': [], 'recovery_time': []
}
for ini, end in zip(true_list, true_list[1:]):
indx_onsets = np.where(aux1[ini:end])[0]
if len(indx_onsets) >= 2: # Valid SCR
start = ini + indx_onsets[0]
finish = end
peaks['start'].append(start)
peaks['end'].append(end)
# Amplitude
segment = gsr_data.phasic[start:finish]
peak_amp = segment.max()
peaks['amp'].append(np.abs(peak_amp - gsr_data.phasic[start]))
# Peak location
peak_loc = np.where(segment == peak_amp)[0][0]
peaks['peak_locs'].append(start + peak_loc)
# Temporal characteristics
peaks['rise_time'].append(peak_loc)
peaks['recovery_time'].append((finish - start) - peak_loc)
return pd.DataFrame.from_dict(peaks)
def compute_tonic_features(gsr_data, fs, seconds, overlap=0.9):
"""
Tonic features with overlapping windows.
Features:
- offset: Linear regression intercept (baseline level)
- slope: Linear regression slope (trend)
- std: Standard deviation (stability)
Args:
overlap: Window overlap fraction (0.9 = 90%)
"""
step = int((1 - overlap) * fs * seconds)
length = fs * seconds
windows = int((len(gsr_data) - length) / step) + 1
tonic = {'offset': [], 'slope': [], 'std': []}
for i in range(windows):
ini = i * step
end = ini + length
# Linear trend
offset, slope = estimate_coefs(
np.arange(0, length),
gsr_data[ini:end]
)
tonic['offset'].append(offset)
tonic['slope'].append(slope)
tonic['std'].append(np.std(gsr_data[ini:end]))
return tonic
Application to Emotion Recognition
Arousal Detection
- SCR frequency ↑: High arousal (excitement, stress)
- SCR amplitude ↑: Intense emotional response
- Tonic level ↑: Sustained arousal state
Cognitive Load
- Tonic slope positive: Increasing mental effort
- SCR rise time ↓: Faster autonomic reaction
- Tonic std low: Stable emotional state
Real-World Application: Affective Robot Storytelling
BIOSIGNALS was the cornerstone of our Affective Robot Story-telling research, where a NAO humanoid robot adapted its narrative delivery based on children's real-time emotional responses detected through physiological signals.
Closed-Loop HRI Experimental Protocol
- Baseline Recording (2 min): Capture resting-state GSR/HRV to establish individual baselines for arousal/valence normalisation
- Story Segments (5 min each): Robot narrates emotional story sections (happy, sad, scary) whilst BIOSIGNALS streams data
- Real-Time Classification (5s windows): Emotion classifier processes HRV+GSR features, emits arousal/valence predictions
- Adaptive Behaviour: Robot modulates voice prosody (pitch, rate), gestures (amplitude, speed), and pacing based on detected emotions
- Event Synchronisation: TCP triggers mark narrative transitions ('segment_start', 'climax', 'resolution') for post-hoc correlation analysis
- Post-Processing: EDF files analysed to correlate physiological changes with specific story moments, validating real-time predictions
"Fusing GSR-derived arousal with HRV-derived valence achieved 74% accuracy in dynamic emotion classification. Crucially, BIOSIGNALS' synchronised event markers revealed that GSR peaks occurred 1-2 seconds after emotionally intense narrative points—a finding that informed our classifier's optimal temporal window of 5 seconds."
Automated Experimental Script
The TCP/IP trigger system enabled fully automated experiments with millisecond-precision event marking:
# Automated multi-trial experimental protocol
from COM.trigger_client import trigger_client
import time
import random
tc = trigger_client('192.168.1.100', 10000)
tc.create_socket()
tc.connect()
stimuli = ['neutral', 'happy', 'sad', 'fear', 'anger']
trials = 20
for trial in range(trials):
# Start recording with baseline
tc.send_msg(b'start')
time.sleep(2) # 2s baseline
# Present randomised stimulus
stimulus = random.choice(stimuli)
tc.send_msg(stimulus.encode())
print(f"Trial {trial}: {stimulus}")
time.sleep(5) # 5s stimulus presentation
# Recovery period
tc.send_msg(b'recovery')
time.sleep(3)
# Stop and auto-save with trial number
tc.send_msg(b'stop')
time.sleep(1) # Inter-trial interval
print("Experiment completed! EDF files saved with timestamps.")
Temporal Precision Analysis
Our analysis of 150+ experimental sessions revealed:
- TCP trigger jitter: Mean 12ms, SD 4ms (acceptable for physiological signals)
- EDF annotation precision: 0.1s (format limitation, sufficient for HRI)
- Emotional response latency: GSR onset 1-2s post-stimulus, HRV changes 3-5s
Technical Implementation: Critical Design Decisions
1. Why PyQt5 + QwtPlot Over Modern Web Frameworks?
In 2018, the natural choice for data visualisation might have been Electron, React, or Plotly Dash. However, for real-time physiological signal processing, PyQt5 offered decisive advantages:
PyQt5 Advantages
- Native performance: Direct GPU rendering, no browser overhead
- Low-latency plotting: QwtPlot achieves 60 FPS at 64 Hz sampling
- Native hardware access: Direct Bluetooth/Serial without sandboxing
- Integrated event loop: Qt signals/slots for thread communication
- Offline reliability: No internet dependency during experiments
Web Framework Limitations
- Rendering lag: Canvas/WebGL redraws lag at high frequencies
- Memory leaks: Long-running sessions degrade performance
- Hardware access: Web Bluetooth/Serial APIs were immature in 2018
- Thread model: Web Workers lack shared memory primitives
- Dependency hell: npm package ecosystem instability
# Precision timer for 64 Hz BVP plotting
self.bvp_timer = QtCore.QTimer()
self.bvp_timer.setTimerType(QtCore.Qt.PreciseTimer)
self.bvp_timer.timeout.connect(self.bvp_update)
self.bvp_timer.start(int((1 / 64) * 1000)) # 15.625 ms
def bvp_update(self):
"""Update BVP plot with minimal latency"""
data = self.dmgs[0].getSamples() # Thread-safe buffer access
self.bvp_curve.setData(data[:, 0], data[:, 1])
self.bio_graph.qwtPlot_bvp.replot() # Native Qt replot
2. EDF+ Format: Why Not HDF5 or Parquet?
| Criterion | EDF+ | HDF5 | CSV/Parquet |
|---|---|---|---|
| Clinical standard | ✓ (ISO/CEN approved) | ✗ | ✗ |
| Toolbox support | ✓ (EEGLAB, FieldTrip, MNE) | △ (requires conversion) | △ (no metadata) |
| Multi-rate channels | ✓ (per-channel sampling) | ✓ | ✗ (single rate) |
| Embedded annotations | ✓ (EDF+ native) | △ (separate dataset) | ✗ |
| File size | △ (no compression) | ✓ (GZIP/LZF) | ✓ (Snappy) |
| Streaming write | ✓ (append mode) | ✓ | ✗ (requires finalisation) |
Decision rationale: For a research tool targeting the neuroscience community, compatibility with established pipelines (EEGLAB, MNE-Python) was paramount. EDF+ provides this whilst maintaining embedded annotations—critical for event-related analysis.
class edf_writter:
def annotation(self, instant, duration, event):
"""Write timestamped annotation to EDF+ file"""
self.file.writeAnnotation(instant, duration, event)
def save_streamData(self):
"""Incremental write during recording"""
for i in range(len(self.data_store)):
self.file.writeSamples(self.data_store[i])
self.file.close()
3. Thread Architecture: Separating Concerns
BIOSIGNALS employs 5 concurrent threads with minimal contention:
- Main GUI thread: PyQt5 event loop, user interactions
- Empatica socket thread: TCP connection to E4 Server, data reception
- Data manager threads (x4): Per-signal buffer management, EDF writing
- Trigger server thread: TCP server for remote commands
- Visualisation timer threads: Per-signal plot updates
class data_manager(QtCore.QThread):
"""Thread-safe per-signal data management"""
def __init__(self, signal, sample_rate):
super().__init__()
self.mutexBuffer = Lock()
self.buffer = RingBuffer(...)
def appendSample(self, sample):
"""Thread-safe insertion"""
self.mutexBuffer.acquire()
try:
self.buffer.append(sample)
self.cur_index += 1
finally:
self.mutexBuffer.release()
def getSamples(self):
"""Thread-safe retrieval for plotting"""
self.mutexBuffer.acquire()
try:
return self.buffer.get()
finally:
self.mutexBuffer.release()
Concurrency Without Deadlocks
By using per-signal mutexes rather than a global lock, we eliminate lock contention between signals. The GUI thread only acquires locks during plot updates (~30ms), whilst data acquisition threads hold locks for <1ms per sample.
Lessons Learned & Future Enhancements
What Worked Exceptionally Well
- ✅ TCP/IP trigger system: Enabled seamless integration with PsychoPy, Unity, ROS, and custom experiment controllers—adopted by 3+ external research groups
- ✅ Modular architecture: MODULES/ directory allowed dynamic loading of custom processing pipelines without modifying core code
- ✅ Open-source release (GPL-3.0): 50+ stars on GitHub, cited in 5+ publications, DOI: 10.5281/zenodo.3759262
- ✅ Real-time HRV processing: 200ms latency for 60s windows enabled closed-loop biofeedback experiments
Trade-offs & Known Limitations
- ⚠️ Hardware dependency: Tightly coupled to Empatica E4 protocol (migration to Polar H10, BITalino, or Shimmer requires refactoring
empatica_client.py) - ⚠️ No cloud synchronisation: Multi-site collaborative studies require manual EDF file sharing (no real-time data federation)
- ⚠️ Limited preprocessing: Artifact removal (motion, electromagnetic interference) performed post-experiment—online filtering would improve classification accuracy
- ⚠️ Windows size trade-off: HRV frequency-domain analysis requires ≥60s windows, limiting temporal resolution for dynamic emotion tracking
If I Were to Rebuild Today (2025 Perspective)
Modern Architecture Proposals
- 🔄 Plugin system via abstract base classes: Define
PhysiologicalDeviceinterface for multi-vendor support (E4, Polar, Muse, Arduino-based DIY sensors) - 🔄 Lab Streaming Layer (LSL) integration: Replace custom TCP protocol with LSL for standardised multi-modal synchronisation (EEG + Biosignals + Motion Capture)
- 🔄 Real-time artifact detection: Implement accelerometer-based motion artifact classifier with automatic correction (wavelet denoising, template matching)
- 🔄 RESTful API + WebSocket server: Enable web-based experiment platforms (PsychoPy3, jsPsych) to control via HTTP alongside legacy TCP
- 🔄 Docker containerisation: Package with all dependencies for reproducible deployments across labs
- 🔄 Machine learning integration: Load pre-trained emotion classifiers (ONNX, TensorFlow Lite) for on-device inference without external servers
Performance Metrics & Validation
Benchmarked on Intel i7-8550U (4 cores, 1.8-4 GHz) with 16GB RAM, Ubuntu 18.04:
| Operation | Latency | Throughput | Notes |
|---|---|---|---|
| Buffer insertion (per sample) | <1 ms | — | O(1) circular buffer |
| GUI update (BVP, 64 Hz) | 15.6 ms | ~64 FPS | QwtPlot native rendering |
| HRV processing (60s window) | ~200 ms | — | Welch PSD + 10 features |
| GSR decomposition (60s) | ~150 ms | — | Savitzky-Golay + SCR detection |
| TCP trigger round-trip (LAN) | <50 ms | — | Measured with 1000 pings |
| EDF write (per sample) | <10 ms | — | Asynchronous I/O thread |
Temporal Precision Validation
We validated temporal accuracy by:
- Hardware test: Sent 5V pulses to E4 whilst recording GSR → detected with 12±4ms jitter
- Cross-modal sync: Compared trigger timestamps in EDF files from simultaneous BIOSIGNALS + EEG recordings → <20ms discrepancy
- Human validation: Participants pressed button whilst watching timestamp display → perceived delay <50ms (imperceptible)
Impact & Scientific Validation
BIOSIGNALS has been released as open-source software with a permanent DOI for academic citation, and directly contributed to multiple peer-reviewed publications in affective computing and human-robot interaction:
💾 BIOSIGNALS Software Release
Repository: github.com/mikelval82/BIOSIGNALS | License: GPL-3.0
Permanent archival on Zenodo ensures long-term accessibility and reproducibility for the research community.
📄 Real-Time Multi-Modal Estimation of Dynamically Evoked Emotions Using EEG, Heart Rate and Galvanic Skin Response
Authors: Mikel Val-Calvo, José Ramón Álvarez-Sánchez, José Manuel Ferrández-Vicente, Eduardo Fernández-Jover
Journal: International Journal of Neural Systems, Vol. 30, No. 4 (2020)
Impact: BIOSIGNALS enabled real-time fusion of GSR, HRV, and EEG for 74% accuracy in dynamic emotion classification. First demonstration of millisecond-precision trigger synchronisation across three modalities in an HRI context.
View Publication📄 Affective Robot Story-telling Human-Robot Interaction: Exploratory Real-time Emotion Estimation
Conference: IEEE RO-MAN 2020 (International Conference on Robot and Human Interactive Communication)
Impact: First adaptive storytelling robot using BIOSIGNALS for closed-loop emotional feedback. Demonstrated 2-second emotional response latency detection enabling truly reactive social robotics.
View Publication🎓 PhD Thesis: Emotional Human-Robot Interaction Using Physiological Signals
Institution: Universidad Nacional de Educación a Distancia (UNED) | Year: 2021
Contribution: Chapter 4 details BIOSIGNALS architecture, validation experiments, and integration within the multi-modal emotion recognition pipeline combining EEG (MULTI_GEERT system), facial expressions, and physiological signals.
Read ThesisCommunity Adoption
Since its 2020 release, BIOSIGNALS has been:
- ⭐ Starred by 50+ researchers on GitHub
- 📚 Cited in 5+ peer-reviewed papers (affective computing, BCI, HRI domains)
- 🏫 Adopted by 3+ research labs for experimental setups beyond the original HRI scope
- 🌍 Downloaded 200+ times from Zenodo archive
Getting Started with BIOSIGNALS
System Requirements
Software Dependencies
Python 3.6+
PyQt5 >= 5.12
pyhrv >= 0.4.0
PythonQwt >= 0.8.0
scipy >= 1.5.0
pandas >= 1.0.0
matplotlib >= 3.0.0
numpy >= 1.18.0
pyEDFlib >= 0.1.20Hardware Requirements
- Empatica E4 wristband (BVP, GSR, TMP, ACC sensors)
- Empatica Server (provided by manufacturer)
- Bluetooth: BLE 4.0+ for E4 connection
- CPU: Quad-core 2 GHz+ (for real-time processing)
- RAM: 8GB minimum (16GB recommended)
- OS: Windows 10/11, Linux (Ubuntu 18.04+), macOS 10.14+
Installation & Quick Start
# 1. Clone repository
git clone https://github.com/mikelval82/BIOSIGNALS.git
cd BIOSIGNALS
# 2. Install dependencies
pip install -r requirements.txt
# 3. Configure Empatica Server (config.py)
E4_server_ADDRESS = 'localhost' # Or remote IP
EMPATICA_PORT = 8000
# 4. Launch application
python BIOSIGNALS_APP_01.py
# GUI Workflow:
# - Set participant ID and output directory
# - Click "Server" to connect to Empatica Server
# - Click "Refresh" to discover E4 devices
# - Select device from dropdown
# - Click "Connect" to start streaming
# - Click "Trigger" to enable remote control (port 10000)
# 5. Remote control from another terminal/script
python
>>> from COM.trigger_client import trigger_client
>>> tc = trigger_client('localhost', 10000)
>>> tc.create_socket()
>>> tc.connect()
>>> tc.send_msg(b'start') # Begin recording
>>> tc.send_msg(b'baseline') # Annotate event
>>> tc.send_msg(b'stop') # Save and increment trial
Important: Empatica E4 Requirement
BIOSIGNALS currently supports Empatica E4 wristbands exclusively. To adapt for other devices:
- Modify
COM/empatica_client.pyto implement your device's protocol - Adjust sampling rates in
constants.py(BVP_SAMPLERATE, GSR_SAMPLERATE, etc.) - Update data parsing logic in
data_manager.py
Community contributions for multi-device support are welcome via pull requests!
Citation & Licence
BibTeX Citation
@software{biosignals2020,
author = {Val Calvo, Mikel},
title = {{BIOSIGNALS: Real-time Physiological Signal
Acquisition System for Emotion Recognition}},
year = 2020,
publisher = {Zenodo},
version = {v1.0},
doi = {10.5281/zenodo.3759262},
url = {https://doi.org/10.5281/zenodo.3759262},
note = {Open-source software for multi-modal
biosignal acquisition with TCP/IP
synchronisation and EDF+ export}
}
@phdthesis{valcalvo2021emotional,
author = {Val Calvo, Mikel},
title = {{Emotional Human-Robot Interaction Using
Physiological Signals}},
school = {Universidad Nacional de Educación a Distancia},
year = {2021},
url = {https://espacio-pre.uned.es/entities/publication/
0e85194e-6187-4e8d-a34c-ca07e5880bd8/full}
}
📜 Licence: GNU General Public Licence v3.0
You are free to:
- ✓ Use commercially and privately
- ✓ Modify and distribute
- ✓ Use for patent claims
- ⚠️ Derivative works must maintain GPL-3.0
- ⚠️ Source code disclosure required for distributions

Resources & Community
Related Open-Source Projects
BIOSIGNALS is part of a larger ecosystem for multi-modal physiological computing:
MULTI_GEERT
EEG Acquisition System
Companion software for OpenBCI, Emotiv, and g.tec amplifiers. Synchronises with BIOSIGNALS via shared TCP trigger server for true multi-modal emotion recognition (EEG + Biosignals + Facial Expressions).
Emotion ML Pipelines
Classification Models
Pre-trained scikit-learn and PyTorch models for arousal/valence classification from HRV+GSR features. Includes feature selection notebooks and cross-validation protocols used in published research.
Additional Resources
- PyHRV Documentation: pyhrv.readthedocs.io — Comprehensive HRV analysis library
- EDF+ Specification: edfplus.info — Official EDF+ format documentation
- HRV Standards: Task Force Guidelines (1996) — Clinical HRV measurement standards
- Empatica E4: support.empatica.com — Device specifications and API docs
Future blog posts will explore MULTI_GEERT architecture, real-time artifact removal with knowledge graphs, and deep learning for neuroprosthetics. Stay tuned!
About the Author
Mikel Val Calvo, PhD
AI Research Scientist specialising in affective computing, neuroprosthetics, and human-robot interaction. Former researcher at Universidad Miguel Hernández de Elche's NeuraViPeR (H2020) project. Currently developing LLM-powered solutions for digital health at LabLENI-UPV.
Join the Discussion
Have questions about implementing physiological signal acquisition for your research? Working on similar affective computing projects? I'd love to hear about your use case and help troubleshoot integration challenges.
Get in Touch